Integrating new statistical frameworks into eDNA survey and analysis at the landscape scale
In recent years, three major innovations have occurred in ecology. (1) The emergence of new statistical methods for analysing community data; (2) the rapid detection of species and whole communities from environmental DNA (eDNA) and bulk-sample DNA; and (3) the wide availability of remotely sensed environmental covariates.
The efficiency gains are such that hundreds or even thousands of species can now be detected and, to an extent, quantified in hundreds or even thousands of samples. Collectively, these three innovations have the potential to relieve the problems of data limitation and analysis that environmental management has been struggling with, opening the way to near-real-time tracking of state and change in biodiversity and its functions and services over whole landscapes.
Principle Investigator Eleni Matechou introduces the project in this short video
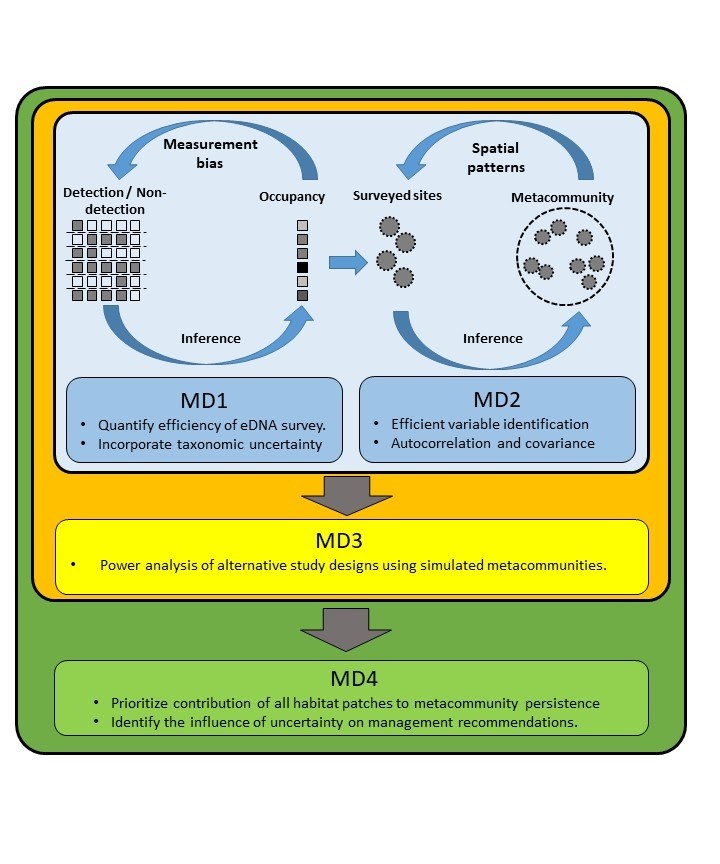
The aim of our project is to develop an integrated statistical framework for DNA-based surveys of biodiversity. The framework will allow the estimation of community compositions and the identification of the landscape characteristics that drive them. We will develop a Bayesian hierarchical model accounting for the probabilistic nature of DNA-based data due to observation error and taxonomic uncertainty and for model uncertainty due to the unknown strength and direction of landscape effects on the system.
We will build sophisticated and efficient algorithms within a Bayesian framework for identifying the important landscape covariates that predict community structure and provide guidelines on optimal allocation of resources in DNA-based surveys for achieving the required power to infer species distributions and to link them to landscape covariates.

Useful links
Project blog: https://blogs.kent.ac.uk/edna/
Rshiny app:
The app accessed on the Shiny server or, for more stable performance, it can be downloaded by going to the Download tab and following the steps outlined to run it on your own machine
Partners website: https://naturespaceuk.com
